|
AbstractThe phosphorylation state of the myosin thick filament-associatedmini-titin, twitchin, regulates catch force maintenance in molluscan smoothmuscle. The full-length cDNA for twitchin from the anterior byssus retractormuscle of the mussel Mytilus was obtained using PCR and 5′rapidamplification of cDNA ends, and its derived amino acid sequence showed a largemolecule (∼530 kDa) with a motif arrangement as follows:(Ig) 11(IgFn 2) 2Ig(Fn) 3Ig(Fn) 2Ig(Fn) 3(Ig) 2(Fn) 2(Ig) 2FnKinase(Ig) 4. Other regions of note include a 79-residue sequencebetween Ig domains 6 and 7 (from the N terminus) in which more than 60% of theresidues are Pro, Glu, Val, or Lys and between the 7th and 8th Ig domains, aDFR XXL motif similar to that thought to be necessary for highaffinity binding of myosin light chain kinase to F-actin. Two majorphosphorylation sites, i.e.
D1 and D2, were located in linker regionsbetween Ig domains 7 and 8 and Ig domains 21 and 22, respectively. Correlationof the phosphorylation state of twitchin, using antibodies specific to D1 andD2, with mechanical properties suggested that phosphorylation of both D1 andD2 is required for relaxation from the catch state. EXPERIMENTAL PROCEDURESMussels—Live specimens of the mussel Mytilusgalloprovincialis for molecular biology experiments and in vitrophosphorylation of isolated twitchin were purchased from the Tokyo CentralWholesale Market.
The ABRM were carefully dissected to avoid any possiblecontamination with visceral parts, immediately frozen in liquid nitrogen, andstored at –80 °C until use. ABRM for mechanical and phosphorylationmeasurements on intact and permeabilized muscles were obtained fromMytilus edulis (Anastasi's Fish Market, Philadelphia, PA) and wereisolated as described previously.5′ -RACE—Total RNA was prepared from ABRMaccording to the method of Chomczynski and Sacchi. First strand cDNA wassynthesized using a 5′-RACE kit (Invitrogen) according to themanufacturer's manual. The sequences of primers used for 5′-RACE areshown in. The startingprimers, TWR1, TWR2, and TWR3, were designed from the sequence of the kinasedomain of Mytilus twitchin described previously. Other primers weredesigned from the sequence determined in the previous 5′-RACEs so thatsequences overlap at least 100 bp. Totally, 43 primers were designed and usedfor cDNA cloning to cover the whole open reading frame.
Each PCR was carriedout as follows: denaturation at 98 °C for 1 s, annealing at 60 °C for5 s, and polymerization at 72 °C for 15 s using a Z- Taq DNApolymerase (TaKaRa). This cycle was repeated 30 times without the finalextension step. The regions amplified by PCR are shown in. The DNA fragmentsamplified by PCR were subcloned into a plasmid vector, pT7 Blue (Novagen), andsequencing was performed by using Dye Deoxy terminator cycle sequencing kitswith a DNA sequencer model 373S (Applied Biosystems). Structural Analysis of Twitchin Motifs—The deduced aminoacid sequence of Mytilus twitchin was analyzed by a Single ModularArchitecture Research Tool (SMART).Mechanical Measurements—ABRM were dissected, andmeasurements of force production were performed under isometric conditionsusing an apparatus that was described previously(,).
Intact ABRM wereincubated in artificial seawater alone or with additionsof acetylcholine (50 μ m) or serotonin (10 μ m).Muscles were permeabilized by incubation in 1% Triton X-100, and the detailedcomposition of the rigor (no ATP), relaxing (1 m m MgATP;pCa 8), and activating (1 m m MgATP; pCa 5)solutions are the same as those used in earlier studies(,). Muscles were frozen inliquid nitrogen while on the mechanical apparatus, pulverized in frozen 0.5n HClO 4, and following thawing and centrifugation, theprotein precipitate was solubilized in SDS sample buffer.SDS-PAGE—SDS-PAGE was performed using buffer conditions ofLaemmli and a totalacrylamide of 5%. Preliminary gels containing the muscle protein samples wererun and stained with Coomassie Brilliant Blue R-250 and subjected to laserdensitometry (Molecular Dynamics) and quantitative analysis (ImageQuantSoftware; Molecular Dynamics).
This allowed adjustment of the protein loadingon subsequent gels so that the same amount of twitchin was added for eachsample used for immunoblotting.Antibodies, Immunoblotting, and Immunodetection—Theantibodies used in this study are those described previously for the D1 and D2sites in twitchin. Theanti-D1 and anti-phospho-D2 antibodies were used at dilutions of 14,000 and500, respectively, for studies on ABRM proteins.Proteins were blotted onto polyvinylidene difluoride and stained with 0.1%Amido Black. Before antibody treatment, the blot was scanned (Umax), and therelative amounts of twitchin in each lane were determined. Immunodetection wasperformed using enhanced chemiluminescence (Amersham Biosciences) and KodakX-OMAT AR imaging film (Eastman Kodak Co.). The film was scanned with a laserdensitometer, and the signal was divided by the relative amount of twitchin onthe blot. Each blot contained samples, as well as five standards ranging fromunphosphorylated twitchin (from a permeabilized ABRM incubated in a rigorsolution for ∼25 min) to maximally phosphorylated twitchin (from apermeabilized muscle incubated in ATP and cAMP). The phosphorylation level inthe sample was determined by comparison to the ECL/protein versusphosphorylation level in the standards.Phosphopeptide Mapping—Isolated twitchin was phosphorylated by thecatalytic subunit of PKA at 25 °C for 10 min in 20 m m Tris-HCl(pH 7.5) containing 0.5 m m dithiothreitol, 2 m mMgCl 2, and 0.5 m m γ- 32PATP.
This wasfollowed by complete digestion with trypsin at 37 °C for 24 h. Theresulting peptides were electrophoresed on a cellulose plate at 1,000 V for 30min using 2.5% formic acid and 7.8% acetic acid as a running buffer. Thinlayer chromatography was carried out for 9 h in a solution containing 37.5%n-butanol, 25% pyridine, and 7.5% acetic acid. After drying, theplate was exposed to x-ray film.Synthetic Peptides—The D1 peptide was RRPSLVDVIPDWPC andrepresented the sequence 1072 to 1084 with a C-terminal Cys residue. The D1site is Ser-1075.
To check on the selectivity of the D1 antibody a peptidecontaining Ser-1011, i.e. RRPSLVDSMKQHLC, was also synthesized. Thisis termed the DX peptide and represented the sequence 1008 to 1020 with aC-terminal Cys residue. These peptides were synthesized and purified togreater than 90% purity by Macromolecular Resources (Ft.
Collins, CO).Both peptides were phosphorylated by PKA at 25 °C for 20 min under thefollowing conditions: 20 m m HEPES (pH 7.2), 2 m mMgCl 2, 0.5 m m ATP, 100 m m KCl, 0.2m m dithiothreitol, 200 μ m peptide, and 0.5 μg/mlcatalytic subunit of PKA (Upstate Biotechnology, Inc.). Assays usingγ- 32PATP indicated that under these conditionsphosphorylation of the peptides reached a maximum at 3 min. The phosphorylatedpeptides in these assay conditions were used directly for dot blots. The twopeptides were equivalent substrates for PKA.Dot Blots—The reactivity of the phosphorylated anddephosphorylated D1 and DX peptides with the affinity-purified antibody to theD1 peptide was monitored using dot blots.
The peptide stock solutions(concentrations determined with the bicinchoninic acid reagent, BCA; Pierce)were prepared in 0.2 mg/ml bovine serum albumin. Samples (10 μl) wereapplied in triplicate to nitrocellulose (Trans-Blot Transfer Medium, 0.2μm; Bio-Rad), air-dried, blocked with 5% non-fat dried milk in 20m m Tris-HCl (pH 7.5), 150 m m NaCl (TBS) for 30 min, andwashed twice for 5 min with TBS. Incubation with primary antibody (D1 antibodyat 14,000× dilution) was for 1 h, followed by two 5-min washes in TBS.The goat anti-rabbit horseradish peroxidase-labeled antibody (Chemicon) at4000× dilution in 1% non-fat dried milk, TBS, was applied for 1 h,followed by two washes for 5 min in TBS. After incubation with the SuperSignalWest Pico Chemiluminescent Substrate (Pierce) for 1.5 min the membrane wasexposed for 5 min to Kodak X-OMAT AR film. The ECL signals were quantitated bydensitometry with Labworks Image Acquisition and Analysis Software and a CDcamera (UVP, Inc.).
The threshold and outline diameter were set at 1% and 65,respectively. The mean integrated optical density from two or three blots wasrecorded (arbitrary units).Mass Spectrometry—Permeabilized ABRM from M.
Eduliswere incubated in a relaxing solution (1 m m MgATP; pCa 8) for 5 min and then an additional 5 min either with or without cAMP(100 μ m). Muscles were frozen, and proteins were subjected toSDS-PAGE. The twitchin-containing band was subjected to in-gel trypticdigestion, and the peptides were analyzed by surface-enhanced laserdesorption/ionization mass spectrometry (PBS II; Ciphergen), in the ProteinChemistry Facility of the Kimmel Cancer Institute at Thomas JeffersonUniversity. RESULTSComplete Sequence of Mytilus Twitchin—Previously a partialsequence of Mytilus twitchin was reported, and it consisted of theC-terminal 878 amino acids. Based on the estimated molecular mass (fromSDS-PAGE) of twitchin of 600 kDa this represented only about one-sixth of thetotal molecule.
This C-terminal fragment contained a kinase domain, four Igmotifs, and one of the sites phosphorylated by PKA, namely the D2 site. In thepresent study, additional sequencing from the kinase domain to the N terminuswas carried out for Mytilus twitchin using fifteen successive roundsof 5′-RACE. This generated a new cDNA sequence of 11.7 kbp spanning theregion from the kinase domain to the 5′-untranslated region, includingthe initiation codon. The new cDNA, when combined with that previouslyreported, contained 16.1 kbp and a coding region of 14.2 kbp. From the latter,the deduced amino acid sequence of full-length twitchin was obtained, and it represented4736 residues with a calculated molecular mass of 526,838. It contains 24 Igand 15 fibronectin type III motifs, in addition to the single kinase domain.The fibronectin type III motifs are well conserved with homologies of 30 to50%. The Ig motifs show lower homology, between 10 and 25%, except for thefive Ig motifs at the N terminus that share about 40% homology.
Other regionsthat should be pointed out include the following: the sequence 790 to 868,between Ig domains 6 and 7, in which more than 60% of the residues are Pro,Glu, Val, or Lys, and an ionic region, 982 to 1009, in which 18 of the 28residues are charged. Phosphorylation Sites—Two phosphopeptides were isolatedpreviously from Mytilus twitchin phosphorylated by PKA.
These were RPSLVDVIPDWP(D1 peptide) and RPSMSPAPEV (D2 peptide). As shown in, the D1 and D2 peptidesequences are 1073–1084 and 4314–4323, respectively and the D1 andD2 sites are Ser-1075 and Ser-4316, respectively. D2 is between the kinasedomain and the C terminus in a linker region connecting Ig motifs 21 and 22,and D1 is in the N-terminal part of the molecule in a linker region between Igmotifs 7 and 8. The D1 and D2 peptides were isolated and sequenced. Attemptsto identify additional sites for PKA by isolating and sequencing thephosphopeptides from twitchin were unsuccessful.
Other putative PKA sitesinclude Ser-1011 (termed DX), Ser-1043, and Ser-1067, with flanking sequencesof RRPSLVD, RKLSRDN, and RRSSMQQ, respectively. In vitrophosphorylation of twitchin with the catalytic subunit of PKA incorporatesapproximately 3 mol P/mol twitchin. A phosphopeptide map of suchphosphorylated twitchin is shown in. It shows two major spots containing similar radioactivities anda minor spot with much less radiolabeled peptide. These data raise thepossibility that incorporation of 2 mol P/mol twitchin result fromphosphorylation of two major sites and that the additional mol P/mol twitchinresults from the sum of phosphorylation of several minor sites. Because the sequencing was done on twitchin from M.galloprovincialis, and the mechanical experiments were performed onM.
Edulis, comparative cDNA sequencing was performed on M.edulis using RT-PCR and primers based on the M.galloprovincialis twitchin sequence. These showed that the derived aminoacid sequences of D1 and D2 in M. Edulis were identical to thesequences of the D1 and D2 M. Galloprovincialis peptides used togenerate the phosphorylation-dependent antibodies. Of note with respect to themass spectrometry experiments described below was a substitution of Val-4326with Ile in M. Edulis twitchin. Although this is not in the regionincluded in the synthetic D2 peptide, it does change the mass of the trypticpeptide containing the D2 phosphorylation site in M.
Theother putative PKA sites near D1 and their flanking regions (RRPSLVD, RKLSRDN,and RRSSMQQ) are also the same in the two species.Because the DX site contained several residues identical to those in the D1peptide ( i.e. RRPSLVD) it was a concern whether the DX peptidecross-reacted with the D1 antibody.
Thus the DX peptide was synthesized andcompared in reactivity to the D1 antibody to the D1 peptide. The sequencesaround the other putative sites were not similar to the D1 peptide and thuswere considered unlikely to cross-react with D1 antibody.Dot Blots—Dot blots were used to examine the sensitivity ofthe D1 antibody with phosphorylated and dephosphorylated D1 and DX peptides.The responses of the D1 antibody over a wide range of peptide amounts areshown in. Theunphosphorylated D1 peptide showed the strongest reaction. Theunphosphorylated DX peptide also reacted, but with a reduced signal comparedwith D1.
Phosphorylation of both peptides considerably reduced their response,and the signal from phosphorylated DX was not detected even at very highpeptide amounts. The responses of the antibody over lower peptide amounts areshown in.Detection of the unphosphorylated D1 peptide was the most sensitive, and areduced response was observed for phosphorylated D1. Both unphosphorylated andphosphorylated DX were not detected over this range. The load of twitchin usedin Western blots was between 0.2 and 1.0 μg, corresponding to about 0.4 to1.9 pmols. The limit of detection in the dot blots for the dephosphorylated D1peptide was between 1 and 2 ng, i.e. About 0.6 to 1.2 pmols.
Thus, atthe low levels of twitchin used on blots in the fiber experiments essentiallyall of the signal detected with the D1 antibody would come from thedephosphorylated D1 site. An important point is that phosphorylated D1 intwitchin would be detected only at concentrations of twitchin exceeding 6pmols, i.e. Considerably above the usual levels of loading.
Inaddition, any signal from the potentially competitive DX peptide( i.e. With the D1 antibody) would not be detected.
4.Sensitivity of the anti-D1 antibody against unphosphorylated andphosphorylated D1 and DX peptides. The integrated optical density of theECL signal is shown as a function of the peptide content in each dot. Dotblots were carried out as described under “ExperimentalProcedures.” Panel A, antibody responses over a range of highpeptide concentrations. Panel B, antibody responses over a range oflow peptide concentrations. Unphosphorylated D1, ♦; phosphorylated D1,.; unphosphorylated DX, ▴; phosphorylated DX, ▪. Phosphorylation of the D1 and D2 Sites in ABRM—We used theanti-D1 antibody and the previously described anti-phospho-D2 antibody to develop an assay forD1 and D2 site phosphorylation in twitchin from intact and permeabilized ABRM.Permeabilized ABRM incubated in a rigor solution for ∼25 min was used asthe source of unphosphorylated twitchin, whereas muscle incubated in ATP andcAMP was the source of maximally phosphorylated twitchin.shows therelative ECL signals from different mixtures of these samples. The anti-D1antibody showed no detectable signal from the phosphorylated sample and analmost linear increase as the fraction of phosphorylated sample was decreased.The anti-phospho-D2 antibody gave ∼8% of maximum signal from theunphosphorylated sample and increased with increasing fraction ofphosphorylated sample.
All phosphorylation measurements of the D1 and D2 sitesusing this technique are reported relative to the maximum phosphorylation ofthe site that is obtained with treatment of the muscle with 100μ m cAMP. 5.Phosphorylation of the D1 and D2 sites of twitchin in intact ABRM. Panel A, standard curve showing the relationship between ECL/proteinand the fraction of phosphorylated sample. The source of unphosphorylatedtwitchin was permeabilized ABRM incubated in a rigor solution for ∼25 min,and the source of maximally phosphorylated twitchin was muscle incubated inATP and cAMP. The intermediate points were mixtures of these samples. ○and dotted line, anti-D1 antibody;. and solid line,anti-phospho D2 antibody.
N = 8 for each. Panel B, designand typical force traces ( solid lines) for determination of D1 and D2phosphorylation in intact ABRM. Muscles were frozen at the end of eachprotocol.
A, catch protocol. Treatment with artificial seawater (ASW)containing acetylcholine (5 × 10 – 5 m) for 1.5 min followed by four ASW washes of 0.5 min each isshown. B, serotonin. Treatment with ASW containing serotonin(10 – 5 m) for 0.5 min and frozen isshown. C, serotonin and then catch. Treatment was as in b,followed by a 15-min ASW wash and then treatment as in a, is shown.d, catch and then serotonin. Treatment was as in a followedby ASW containing serotonin for 0.5 min.
Panel C, D1 ( openbars) and D2 ( striped bars) site phosphorylation for the designsshown in panel B. Data are mean ± S.E. Twitchin from permeabilized ABRM at pCa 8 with and withoutcAMP was subjected to trypsin digestion and surface-enhanced laserdesorption/ionization mass spectrometry.shows that in theabsence of cAMP there is a peptide with a mass (M+H +)corresponding to the unphosphorylated D1 tryptic peptide (2030.4).
CAMPtreatment of the muscle results in the loss of this peptide and the appearanceof a peptide 80 dalton larger, as expected from phosphorylation of the D1site. The almost complete loss of the signal from the unphosphorylated D1peptide suggests nearly 100% phosphorylation of the D1 site. Twitchin from thesame sample also showed no reaction with the D1 antibody confirming theabsence of an interaction of the antibody with the phosphorylated D1 site.
Thesmall amount of phosphorylated peptide in the mass spectrum from the samplewithout cAMP is consistent with the 11% phosphorylation of D1 determined onthe same sample using the above described antibody technique. The smallersignal in the mass spectra from the phosphopeptide compared with theunphosphorylated peptide is not unusual. The only other instanceof a cAMP-mediated decrease in one peak and an increase in another at 80dalton higher M+H + is 2062 to 2142. This does not match anyexpected unmodified tryptic peptide but might result from dihydroxylation ofthe D1 peptide. The unphosphorylated tryptic D2 peptide (1964.3) was notobserved in either spectrum, but the phosphorylated sample showed a decreasein a peptide of m/ z 1978, which might represent a methylatedD2 tryptic peptide. Peaks corresponding to tryptic peptides containingputative phosphorylation sites at Ser-1011, Ser-1043, and Ser-1067 were notobserved in the unphosphorylated sample, so possible phosphorylation of thesesites could not be determined by this method.
6.Surface-enhanced laser desorption/ionization mass spectra of trypticdigests of twitchin from permeabilized ABRM either treated with cAMP ( redline) or not ( green line). Also shown is the differencespectrum ( black line). Peaks A and B show acAMP-mediated decrease in signal from a peptide with mass M+H +corresponding to the unphosphorylated D1 tryptic peptide (2030.4) and theappearance of a peptide with 80 dalton larger mass as expected fromphosphorylation. Uno nessuno centomila ebooking. The only other instance of a cAMP-mediated decrease in onepeak and an increase in another at 80 dalton higher mass is 2062 ( peakC) to 2142 ( peak D).
These may represent the unphosphorylatedand phosphorylated forms of a dihydroxylated D1 peptide. The unphosphorylatedtryptic D2 peptide (1964.3) was not observed in either spectrum, but thephosphorylated sample did show a decrease in a peptide of mass 1978 dalton( peak E), which might represent a methylated D2 tryptic peptide. Intact Muscles—The four experimental designs used todetermine the state of D1 and D2 phosphorylation in intact ABRM underdifferent mechanical conditions are shown in.
Multiplesuccessive activations with acetylcholine were used to maximize catch forcemaintenance , whereastreatment with serotonin results in release of catch force.shows that D2phosphorylation is quite low in catch and high in the presence of serotoninwhen catch force is released. Interestingly, D1 site phosphorylation is about40% in catch and increases to almost 100% in serotonin. The increase inphosphorylation of both D1 and D2 in serotonin is reversed when the muscle issubsequently put into catch.Permeabilized Muscles—To obtain a better understanding ofthe relationships between the phosphorylation of D1 and D2 and the mechanicalresponse of the muscle, experiments were performed on permeabilized ABRM inwhich it was possible to vary cAMP and the degree of twitchinphosphorylation. Shows the design in which muscles were activated inpCa 5 and then switched to pCa 8. In the absence ofcAMP, high catch force is maintained, but as cAMP is increased, the catchforce maintenance is decreased. Shows the force remaining 2 min after transfer topCa 8 (a measure of the catch force), as well as D1 and D2phosphorylation as a function of cAMP. D1 phosphorylation is ∼30% evenin the absence of cAMP and increases to almost 100% as catch force isdecreased.
D2 phosphorylation is very low in the absence of cAMP and increasesto more than 80% as cAMP increases and catch force dissipates.shows therelationships of catch force to D1 and D2 phosphorylation in individualmuscles. D1 phosphorylations from 0 to 50% appear to have little effect oncatch, but higher values are associated with release of catch force. On theother hand, D2 phosphorylation and force show a very different relationship inthat small increases in phosphorylation are associated with changes in catchforce maintenance. 7.Phosphorylation of the D1 and D2 sites of twitchin at different cAMPin permeabilized ABRM.
Panel A, experimental design and typicalforce traces at different cAMP. Muscles were transferred from pCa 8 to pCa 5 for 5 min and then back to pCa 8 for 2min and frozen. Panel B, D1 (○ and dashed line) and D2(.
and solid line) phosphorylation as a function of cAMP. Alsoshown is the fraction of the pCa 5 force remaining at 2 min aftersubsequent transfer to pCa 8 (▴ and dotted line).Data are mean ± S.E.
With n = 3–5 except for the 100μ m points, which are the means of two measurements. PanelC, fraction of force remaining as a function of D1 (○ and dashedline) and D2 (. and solid line) phosphorylation forindividual muscles treated with different cAMP. DISCUSSIONPrimary Structure of Twitchin—The motif arrangements fortwitchin from Mytilus and Caenorhabditis elegans are quitesimilar and are(Ig) 11(IgFn 2) 2Ig(Fn) 3Ig(Fn) 2Ig(Fn) 3(Ig) 2(Fn) 2(Ig) 2FnKinase(Ig) 4for Mytilus and(Ig) 8(IgFn 2) 10Ig(Fn) 3Ig(Fn) 2Ig(Fn) 3(Ig) 2(Fn) 2(Ig) 2FnKinase(Ig) 5for C.
The major difference is that following a series of Igdomains at the N terminus, C. Elegans has ten IgFn 2repeats compared with only two in the Mytilus protein (molecular mass∼750 versus ∼530 kDa, respectively). The Align program atGenestream shows 27.7% identical residues. The partial sequence that isavailable from the kinase domain region of Aplysia twitchin showshigh homology with both Mytilus and C. Elegans proteins. The structuralsimilarity of these molecules from different species suggests that theyperform similar functions.Twitchin, the product of the unc-22 gene in C.
Elegans,was so named, because mutations of the protein resulted in a constanttwitching of the body wall muscle. Extragenic dominant suppressors of theunc-22 mutant phenotype were localized as mutations in the headregion of myosin, and the suppressor phenotypes showed slower than normalmovement.
These resultsraised the possibility that twitchin may interact with the myosin head andthat a mutation that slows myosin cross-bridge cycling rate might reverse theeffects of twitchin mutation. The twitching phenotype may thus result fromremoval of a mechanism that slows myosin cycling rate. This would result infaster contraction and relaxation cycles associated with each activation ofthe muscle in the absence of twitchin. Studies of the mechanical effects oftwitchin phosphorylation support the idea that twitchin controls the kineticsof myosin interaction with actin. Phosphorylation of twitchin inAplysia and inMytilus ABRM (see and Refs., and) is associated with anincreased rate of relaxation.
In the ABRM at low Ca 2 +,myosin detaches very slowly from actin when twitchin is unphosphorylated butdetaches very rapidly when twitchin is phosphorylated. Similarly, studies onnative and synthetic thick filaments from byssus retractor muscles ofMytilus showed that at low calcium the presence of twitchin promotedactin-myosin binding, but only under conditions expected to result inunphosphorylated twitchin. Therefore, a primaryrole of twitchin seems to be to slow the detachment of myosin from actin underlow intracellular Ca 2 +. Phosphorylation prevents thiseffect and seems to mimic the absence of twitchin in studies on C.elegans and on isolated and synthetic Mytilus thickfilaments.The sequence of Mytilus twitchin includes a short PEVK-rich domainbetween Ig domains 6 and 7. The PEVK region of titin overlaps the I-band invertebrate muscles and is thought to contribute to the elastic properties ofthe resting muscle.
Thissequence of 79 amino acids in twitchin is shorter and has a lower frequency ofPEVK residues than titin from vertebrate striated muscle, and its function isunknown. Smooth muscle myosin light chain kinase (MLCK), which is related totwitchin, also contains a PEVK region, and, as in the case of twitchin, itsfunction in the molecule is not clear.
In addition,Mytilus twitchin shares with MLCK the motif (DFR XXL) in theregion between the seventh and eighth Ig domains that also contains the D1phosphorylation site. There are three or five of these motifs in smooth muscleor non-muscle MLCK, and they are thought to be necessary for the high affinitybinding of MLCK to F-actin(–).Twitchin from C. Elegans also contains a DFR XXL motif in thesame area of the molecule (between the fourth and fifth Ig domain).Interestingly, the PEVK domain from cardiac muscle titin also has been shownrecently to interact withactin. The presence of these motifs in twitchin raises the possibility thattwitchin binds to actin, but there was little evidence for this incosedimentation experiments on Mytilus twitchin and F-actin.Site-specific Phosphorylation of Twitchin—The increase inphosphorylation of twitchin and associated relaxation of catch force iscomplete within 30 s after addition of cAMP to permeabilized ABRM and additionof serotonin to intact muscles. In vitrophosphorylation of twitchin with the catalytic subunit of PKA incorporatesapproximately 3 mol P/mol twitchin, but based on the kinetics ofphosphorylation two ofthese sites are preferred.
Because the in vivo phosphorylationtransients also are rapid, these two sites can be considered as the majorphysiological PKA sites. In addition, a phospho-peptide map of invitro phosphorylated twitchin showed two major spots and a minor spot.The results from phosphorylation-dependent antibodies show that two siteswhose phosphorylation increases with relaxation of catch are D1 and D2, andthese are located in the N- and C-terminal regions of the molecule,respectively. Failure to identify other sites could be because of amultiplicity of sites ( i.e. Ser-1011, Ser-1043, and Ser-1067 arecandidates) and/or low stoichiometries of phosphorylation. The potentialrole(s) of phosphorylation at other sites is not known, but based on thepresent results it is clear that phosphorylation transients of D1 and D2 arecompatible with the observed mechanical properties. In this regard, the use ofphosphorylation-dependent antibodies to D1 and D2 was valuable for comparisonof mechanical response to phosphorylation at the D1 or D2 sites.In both intact and permeabilized muscles, the D1 site can be 40 to 50%phosphorylated when catch force is well maintained (see Figs.and), and there is no change inrate of relaxation of force when D1 phosphorylation varies from 0 to 50% inpermeabilized muscles.
This strongly suggests that phosphorylation of D1 aloneis not sufficient to facilitate rapid detachment of myosin from actin at lowcalcium concentrations. The relatively high level of D1 phosphorylationcompared with D2 phosphorylation in catch in the intact muscles and at lowconcentrations of cAMP in permeabilized muscles indicates differentsensitivities of the sites to protein kinase A and/or phosphatase activity.Isolated twitchin shows an endogenous kinase activity, and this is blocked bythe peptide inhibitor of PKA. This phosphorylation wasattributed to the copurification with twitchin of the PKA catalytic subunit orunregulated holoenzyme.
The basal 50% phosphorylation of D1 in intact musclesand the apparent cAMP-independent partial phosphorylation in permeabilizedmuscle may reflect a preferential binding of an unregulatedholoenzyme/catalytic subunit to the D1 location. It could also reflect bindingof another kinase, although this is thought to be unlikely because of theinhibition of endogenous kinase activity by PKA inhibitors.
Even if another kinasecan partially phosphorylate the D1 site, full phosphorylation of D1 appears torequire activation of PKA.Relaxation of catch force only occurs when D2 phosphorylation increases,and this occurs only when D1 phosphorylation is relatively high. Thephosphorylation of both of these sites suggests that a phosphorylation-inducedchange in interaction at both ends of twitchin is necessary for the observedregulation of cross-bridge detachment rate. The long length of twitchin(∼0.2 μ) raisesthe possibility of interactions with multiple myosin molecules, and thisprovides an obvious mechanism by which phosphorylation-induced changes at theends of twitchin are transmitted to many myosin cross-bridges.Comparison to Cardiac Myosin-binding Protein C—Another thickfilament-associated protein that is thought to modulate actin-myosininteraction is myosin-binding protein C (MyBP-C), and it is of interest tocompare what is known about the structural and functional aspects of thisprotein to twitchin. Although the exact role of MyBP-C in cardiac muscle isnot certain, various reports suggest that it regulates force output and/orvelocity of shortening(–)and that phosphorylation of MyBP-C mediates these effects.
Isolated MyBP-C isphosphorylated at four sites. Three of these can be phosphorylated by PKA andare located between the second and third Ig domains from the N terminus,whereas the other, which is phosphorylated by PKC, lies between the Fn and Igdomains at the C terminus of the molecule. Phosphorylation of theN-terminal sites is thought to regulate the mechanical effect of MyBP-C. Indeed, this portion ofthe protein shows phosphorylation-dependent binding to myosin S2.
MyBP-C also has a myosinrod binding site located in the Ig domain at the C terminus(,).shows thatMytilus twitchin has some structural similarities to the cardiacisoform of MyBP-C. The Ig/Fn domain arrangement in MyBP-C is similar to theportion of twitchin near the D1 phosphorylation site. Both proteins havephosphorylation sites near each end of the molecule, and the N-terminalregulatory phosphorylation sites are followed by similar arrangements ofimmunoglobulin and fibronectin motifs (Ig 4Fn 2IgFnand Ig 5Fn 2IgFn in MyBP-C and twitchin,respectively). The interdomain region containing the D1 phosphorylation siteis aligned with the region in MyBP-C containing the phosphorylation-dependentmyosin S2 binding site. These regions of the two molecules do not showsignificant sequence homologies. There is, however, a highly charged areabetween residues 983 and 1009 in twitchin that contains 10 positive and 8negative amino acids, and this charge distribution is similar to an N-terminalportion of caldesmon that has been shown to bind to the S2 region of smoothmuscle myosin(–).Another finding is that there is about 40% sequence identity and 55% homologybetween the Ig domain that is C-terminal to the D2 phosphorylation site intwitchin and the myosin rod binding Ig domain in MyBP-C.
Twitchin and MyBP-Cmay share a structural basis of function that includes each end of themolecule interacting with a different region of myosin with some of theseinteractions modulated by phosphorylation of the protein.
DescriptionOriginal benchmark for HEPA filtration and sterilizationAdvanced incubator design for high value cellsAdvanced large capacity for high value cells and high throughput labsOptimal choice for large scale or GMP applicationsCapacity184L (6.5 cu. Ft.)165L (5.8 cu. Ft.)255L (9.0 cu.
Ft.)232L (8.2 cu. Ft.)322L (11.4 cu.
Ft)Chamber typeDirect heatDirect heatDirect heatDirect heatDoor swingLeft hingedLeft hingedLeft hingedRight hingedChoose interior configuration140°C sterilizationSteri-Run (180°C)Steri-Run (180°C)140°C sterilizationFan assistedTHRIVETHRIVEFan assistedTC or IRTC180 or IR180SiTC180 or IR180SiIR sensorN/AOptional 1–21%or 5–90%Optional 1–21%or 5–90%N/ARemovable water panCovered integrated reservoir with easy drainingCovered integrated reservoir with easy drainingActive control with external reservoirEnviro-ScaniCAN touchscreeniCAN touchscreenEnviro-Scan. DescriptionOriginal benchmark for HEPA filtration and sterilizationAdvanced incubator design for high value cellsAdvanced large capacity for high value cells and high throughput labsOptimal choice for large scale or GMP applicationsCapacity184L (6.5 cu. Ft.)165L (5.8 cu.
Ft.)255L (9.0 cu. Ft.)232L (8.2 cu.
Ft.)322L (11.4 cu. Ft)Chamber typeDirect heatDirect heatDirect heatDirect heatDoor swingLeft hingedLeft hingedLeft hingedRight hingedChoose interior configuration140°C sterilizationSteri-Run (180°C)Steri-Run (180°C)140°C sterilizationFan assistedTHRIVETHRIVEFan assistedTC or IRTC180 or IR180SiTC180 or IR180SiIR sensorN/AOptional 1–21%or 5–90%Optional 1–21%or 5–90%N/ARemovable water panCovered integrated reservoir with easy drainingCovered integrated reservoir with easy drainingActive control with external reservoirEnviro-ScaniCAN touchscreeniCAN touchscreenEnviro-Scan. On-demand high temperature sterilization cycles for easy cleaning. Our designs are independently tested to meet U.S. And EU pharmacopeia standards.
Automatically radiates heat uniformly to all interior surfaces and quickly returns to selected operating conditions. Simplifies cleaning without the need of autoclaving and eliminates variability in manual cleaning. Independent third-party tests prove elimination of biological contaminants including fungi, spores, mycoplasma, and other bacteria. Avoids physical constraints of UV germicidal lamps and ongoing costs, handling and storage of potentially toxic germicides.
Thermo Scientific ContraCon Sterilization Cycle140°C Sterilization CycleThermo Scientific Steri-Run Sterilization CycleTemperature type90°C moist heat cycle140°C dry heat cycle180°C dry heat cycleDescriptionPush button simplicity without removal of sensors and components.Overnight sterilization cycle with extended time at 140°C to achieve sterilization.The new standard, an overnight cycle that reaches 180°C on all chamber surfaces.Duration of sterilization cycle24–25 hours14 hours12 hoursSterility assurance level (SAL)6 log12 log12 logIndependent validation✓✓✓Learn more:Learn more:Learn more. The variable speed fan precisely controls the airflow to provide leading edge recovery. The air stream first travels over the direct-heated covered, integrated water reservoir, which results in 50% faster humidity recovery than a standard water pan design. The in-line HEPA filter cleans the airstream of all sizes of particles and microorganisms, protecting cultures from contamination.
The air then passes over the in-chamber sensors, which react and condition the air before gently circulating it among the cultures. Thermal conductivity (TC) CO 2 sensor. Our most commonly requested CO 2 sensor due to its reliability and longevity of operation. Measures electrical resistance according to thermal conductivity of gas passing over the sensor. Low maintenance, stable, and long life because it has no consuming element such as an incandescent bulb. High temperature resistance—does not need to be removed during sterilization cycles. NEW TC180 CO 2 sensor incorporates humidity compensation for enhanced stability and recovery (available in the Heracell VIOS and Forma Steri-Cycle CO 2 incubators).
Infrared (IR) CO 2 sensor. Ideal for sensitive applications and remote monitoring or when frequent access to incubator is required. Determines the CO 2 concentration based on its IR light absorption at the characteristic wavelength of 4.3 µm.
Download Free Forma Scientific Model 2095 Manual Muscles
Operates independent of humidity and temperature. NEW IR180Si replaces the traditional incandescent IR light source with silicone MEMS emitter technology for improved stability and service life (available in the Heracell VIOS and Forma Steri-Cycle CO 2 incubators).
1Department of Cardiological Sciences, St. George’s Hospital Medical School, London, United Kingdom 2Department of Medicine and Geriatrics, Kochi Medical School, Kochi, Japan 3Departamento de Cardiologia, Clinica Medellin, Medellin, ColombiaAddress correspondence to: J. Mogensen, Department of Cardiological Sciences, St.
George’s Hospital Medical School, Cranmer Terrace, Tooting, SW17 0RE, London, United Kingdom. Phone: 2002; Fax: 0944; E-mail:.Find articles byMogensen, J.in:. 1Department of Cardiological Sciences, St. George’s Hospital Medical School, London, United Kingdom 2Department of Medicine and Geriatrics, Kochi Medical School, Kochi, Japan 3Departamento de Cardiologia, Clinica Medellin, Medellin, ColombiaAddress correspondence to: J. Mogensen, Department of Cardiological Sciences, St. George’s Hospital Medical School, Cranmer Terrace, Tooting, SW17 0RE, London, United Kingdom.
Phone: 2002; Fax: 0944; E-mail:.Find articles byKubo, T.in:. 1Department of Cardiological Sciences, St.
George’s Hospital Medical School, London, United Kingdom 2Department of Medicine and Geriatrics, Kochi Medical School, Kochi, Japan 3Departamento de Cardiologia, Clinica Medellin, Medellin, ColombiaAddress correspondence to: J. Mogensen, Department of Cardiological Sciences, St. George’s Hospital Medical School, Cranmer Terrace, Tooting, SW17 0RE, London, United Kingdom.
Phone: 2002; Fax: 0944; E-mail:.Find articles byDuque, M.in:. 1Department of Cardiological Sciences, St. George’s Hospital Medical School, London, United Kingdom 2Department of Medicine and Geriatrics, Kochi Medical School, Kochi, Japan 3Departamento de Cardiologia, Clinica Medellin, Medellin, ColombiaAddress correspondence to: J.
Mogensen, Department of Cardiological Sciences, St. George’s Hospital Medical School, Cranmer Terrace, Tooting, SW17 0RE, London, United Kingdom. Phone: 2002; Fax: 0944; E-mail:.Find articles byUribe, W.in:.
1Department of Cardiological Sciences, St. George’s Hospital Medical School, London, United Kingdom 2Department of Medicine and Geriatrics, Kochi Medical School, Kochi, Japan 3Departamento de Cardiologia, Clinica Medellin, Medellin, ColombiaAddress correspondence to: J. Mogensen, Department of Cardiological Sciences, St. George’s Hospital Medical School, Cranmer Terrace, Tooting, SW17 0RE, London, United Kingdom. Phone: 2002; Fax: 0944; E-mail:.Find articles byShaw, A.in:. 1Department of Cardiological Sciences, St.
George’s Hospital Medical School, London, United Kingdom 2Department of Medicine and Geriatrics, Kochi Medical School, Kochi, Japan 3Departamento de Cardiologia, Clinica Medellin, Medellin, ColombiaAddress correspondence to: J. Mogensen, Department of Cardiological Sciences, St.
George’s Hospital Medical School, Cranmer Terrace, Tooting, SW17 0RE, London, United Kingdom. Phone: 2002; Fax: 0944; E-mail:.Find articles byMurphy, R.in:.
1Department of Cardiological Sciences, St. George’s Hospital Medical School, London, United Kingdom 2Department of Medicine and Geriatrics, Kochi Medical School, Kochi, Japan 3Departamento de Cardiologia, Clinica Medellin, Medellin, ColombiaAddress correspondence to: J.
Mogensen, Department of Cardiological Sciences, St. George’s Hospital Medical School, Cranmer Terrace, Tooting, SW17 0RE, London, United Kingdom. Phone: 2002; Fax: 0944; E-mail:.Find articles byGimeno, J.in:. 1Department of Cardiological Sciences, St. George’s Hospital Medical School, London, United Kingdom 2Department of Medicine and Geriatrics, Kochi Medical School, Kochi, Japan 3Departamento de Cardiologia, Clinica Medellin, Medellin, ColombiaAddress correspondence to: J. Mogensen, Department of Cardiological Sciences, St.
George’s Hospital Medical School, Cranmer Terrace, Tooting, SW17 0RE, London, United Kingdom. Phone: 2002; Fax: 0944; E-mail:.Find articles byElliott, P.in:. 1Department of Cardiological Sciences, St. George’s Hospital Medical School, London, United Kingdom 2Department of Medicine and Geriatrics, Kochi Medical School, Kochi, Japan 3Departamento de Cardiologia, Clinica Medellin, Medellin, ColombiaAddress correspondence to: J.
Mogensen, Department of Cardiological Sciences, St. George’s Hospital Medical School, Cranmer Terrace, Tooting, SW17 0RE, London, United Kingdom. Phone: 2002; Fax: 0944; E-mail:.Find articles byMcKenna, W.in: First published January 15, 2003. Restrictive cardiomyopathy (RCM) is an uncommon heart muscle disorder characterized by impaired filling of the ventricles with reduced volume in the presence of normal or near normal wall thickness and systolic function. The disease may be associated with systemic disease but is most often idiopathic. We recognized a large family in which individuals were affected by either idiopathic RCM or hypertrophic cardiomyopathy (HCM). Linkage analysis to selected sarcomeric contractile protein genes identified cardiac troponin I (TNNI3) as the likely disease gene.
Subsequent mutation analysis revealed a novel missense mutation, which cosegregated with the disease in the family (lod score: 4.8). To determine if idiopathic RCM is part of the clinical expression of TNNI3 mutations, genetic investigations of the gene were performed in an additional nine unrelated RCM patients with restrictive filling patterns, bi-atrial dilatation, normal systolic function, and normal wall thickness. TNNI3 mutations were identified in six of these nine RCM patients. Two of the mutations identified in young individuals were de novo mutations. All mutations appeared in conserved and functionally important domains of the gene. Restrictive cardiomyopathy (RCM) is a myocardial disease characterized by impaired ventricular filling and reduced diastolic volume in the presence of normal systolic function and normal or near normal myocardial thickness ( ).
The disease is characterized by symptoms of progressive left- and right-sided heart failure. The overall prognosis is poor, especially when onset is in childhood, and patients often require cardiac transplantation (, ). Although several inherited and acquired disorders may cause RCM, many cases remain idiopathic (, ). Familial RCM has been reported, but whether a distinct genetic entity exists remains uncertain ( ).A child and his mother were diagnosed with idiopathic RCM in our cardiomyopathy clinic. Their family history revealed that 12 individuals died suddenly and that several surviving relatives had symptoms of heart disease. Clinical investigation of the family (H640) identified another individual with RCM and nine individuals with hypertrophic cardiomyopathy (HCM).
Genetic investigations identified a disease-causing mutation within a highly conserved region of the cardiac troponin I gene ( TNNI3). To test the hypothesis that RCM is part of the clinical expression of TNNI3 mutations, we performed genetic investigations of the gene in nine additional idiopathic RCM patients. This paper presents the results of clinical and genetic investigations in ten unrelated patients diagnosed with idiopathic RCM and their relatives, including family H640 mentioned above.
This study was approved by the local research ethics committee, and informed consent was obtained from all participants.Patients. Clinical investigations of the initial family (H640) comprised the proband and 32 of his surviving relatives. The remaining study cohort consisted of nine additional unrelated patients diagnosed with idiopathic RCM over a 6-year period. Relatives of these RCM patients were also invited to participate in the study. All patients and those relatives who agreed to participate underwent physical examination, 12-lead ECG, and transthoracic two-dimensional echocardiography and Doppler studies.
Selected patients underwent cardiac catheterization and right ventricular biopsy (H417, H906, H816, H974, H38).Echocardiography. Standard views for M-mode and two-dimensional studies were obtained. End-diastolic left ventricular wall thickness was recorded at the level of the mitral valve and papillary muscle in the anterior and posterior septum and the lateral and posterior wall using short-axis two-dimensional images. Anterior and posterior septal thickness at the apex was assessed from apical four-chamber and parasternal short-axis views. The maximum left ventricular wall thickness was defined as the maximal measurement recorded in any of the myocardial segments studied. Left ventricular wall thickness and left ventricular end-diastolic dimension was adjusted for age and body surface area in RCM patients (Tables and ) (, ).
Septal/posterior wall ratio was obtained from measurements in the parasternal long-axis view, the short-axis view at the mitral valve level, or the short-axis view at the papillary muscle level. Table reports the ratio that was the greater of the two. Left ventricle in- and outflow velocities were determined using continuous and pulse wave Doppler echocardiography. Clinical data of patients diagnosed with idiopathic RCMRestrictive cardiomyopathy. Individuals were diagnosed with RCM in accordance with previous established criteria when echocardiography revealed the following (, ): Doppler measurements consistent with restrictive left ventricular filling pattern (increased ratio of early diastolic filling E to atrial filling A ≥ 2; decreased deceleration time DT ≤ 150 ms; decreased isovolumic relaxation time IVRT ≤ 70 ms); reduced or low-normal left ventricular end diastolic dimension (LVED); normal or near normal left ventricular wall thickness and systolic function.Hypertrophic cardiomyopathy. Individuals were diagnosed with HCM when echocardiography identified unexplained left ventricle hypertrophy greater than or equal to 13 mm, or they fulfilled proposed diagnostic criteria for HCM within the context of familial disease, i.e., echocardiographic criteria and/or one of the following ECG abnormalities (, ): left ventricular hypertrophy (Romhilt-Estes score ≥4); Q-waves (duration 0.04 sec and/or a depth 1/4 of ensuring R wave in at least two leads); and marked repolarization abnormalities (T-wave inversion in at least two leads).DNA extraction. Genomic DNA was extracted from peripheral blood samples using a kit from QIAGEN Inc.
51306; West Sussex, United Kingdom).Linkage, haplotype, and paternity analysis. Linkage analysis of recognized HCM genes in family H640 were carried out using microsatellite markers and the LINKAGE 5.1 package (as described previously ( ). Haplotype analysis of pedigree H816 and H805 was performed using microsatellite markers defining the TNNI3 locus (D19S254, D19S210, D19S877, and D19S887) ( ). Paternity analysis was performed in pedigree H417 and H906 using 13 microsatellite markers (D1S1656, D3S1578, D11S1344, D7S483, D12S1583, MyoI-CA, MyoII-CA, HTMα-CA, D15S1036, D15S974, D15S153, D19S926, D19S877, and D19S887).Mutation analysis.
The eight protein-encoding exons of TNNI3 were amplified using intronic primers designed according to the genomic sequence of the gene using the OLIGO 4.1 primer analysis software (MedProbe AS, Oslo, Norway) ( ). Primer sequences and PCR conditions are available upon request.PCR products were purified prior to cycle sequencing using a kit from Amersham Pharmacia Biotech (no. 25-6902-01; Amersham, United Kingdom) and cycle sequencing was performed using a BigDye Terminator Cycle Sequencing Kit from Applied Biosystems Inc. 4314414; Foster City, California, USA). The sequences were analyzed on an ABI Prism 3100 Sequencer in accordance with the manual of the manufacturer.When a mutation was identified, a second blood sample from the affected patient was reanalyzed to confirm the finding by direct sequencing and conventional SSCP analysis (standard 10% polyacrylamide gel electrophoresis in 1×TBE buffer at room temperature and postelectrophoresis silver staining).
Three of the mutations identified were predicted to change restriction enzyme sites according to computer analysis using Webcutter software (These findings were confirmed by PCR amplification of the relevant exon restriction enzyme digest, followed by size fractionation using either 3% agarose or 12% PAGE gel electrophoresis. Mutation Asp190Gly in pedigree H640 abolished a BsrI site, mutation Arg192His in pedigree H417 abolished a HhaI site, and mutation Arg145Trp in pedigree H816 and H805 created an AciI site. Clinical investigations of family H640.
The proband of family H640, individual IV-10 (Figure, Table ), had onset of heart failure symptoms at the age of 11 years. Echocardiography revealed severe bi-atrial enlargement accompanied by a restrictive filling pattern, normal systolic function, and normal left ventricular wall thickness. His ECG showed prominent P-waves in all leads consistent with bi-atrial enlargement. His mother (III-17) had symptoms of heart failure from the age of 29 years. Her investigations also demonstrated restrictive left ventricular physiology with normal wall thickness measurements, except for localized apical thickening (14 mm).
Pedigree drawings of restrictive cardiomyopathy families affected by cardiac troponin I mutations. Squares indicate male family members; circles, female family members; symbols with slash, deceased individuals; open symbols, unaffected individuals; filled symbols, individuals affected by RCM (indicated by arrows) or HCM; checkered symbols, individuals who died suddenly; question mark, unknown clinical status.
Plus sign indicates presence of mutation, whereas minus sign indicates absence of mutation. Generation II of family H640 consisted of a total of 13 siblings, of which 9 were available for this investigation.
Four generation-II siblings were obligate carriers of the mutation (included in the pedigree drawing with their offspring), while five remaining siblings were shown not to carry the mutation. Two of the obligate carriers (III-7, II-3) died suddenly at the age of 40. The remaining obligate carriers (II-1, II-2, II-4), reached the age of 63, 59, and 51 years, respectively. Ten sudden deaths appeared in young individuals who had a 50% chance of having inherited the disease (IV-1/2/4; III-18/20/21/22/23/26/27). Their average age of death was 25 years (16–33 years), and most had symptoms of cardiac disease prior to their death.
Individual IV-13 was shown to be a healthy carrier of the mutation at the age of 5 years.The family history revealed several relatives with symptoms of heart disease and nine individuals who died suddenly at less than 30 years of age. A cousin of the proband, individual IV-3, had already been diagnosed previously with restrictive cardiomyopathy following echocardiography and ECG-recording (Table ).Nine living relatives within pedigree H640 fulfilled diagnostic criteria for HCM within the context of familial disease (, ). Six of these individuals (III-1, III-3, IV-5, III-6, III-11, III-15) had left ventricular wall hypertrophy of 13–20 mm, including three individuals with a left atrial size of more than 50 mm (III-1, III-6, III-15). The diastolic filling patterns of these individuals were mixed, showing features of restriction as well as impaired diastolic relaxation with a shortened DT, but normal or prolonged IVRT, and diminished to normal E/A ratio in the majority.
Editorial Reviews. From Booklist. It's an irony so commonplace it's become almost trite: despite the information superhighway, despite a world of knowledge at their fingertips, the younger. Bauerlein's simple but jarring thesis is that technology and the digital culture it has created are not broadening the horizon of the younger generation; they are. In his book, The Dumbest Generation, Mark Bauerlein (2008) addresses some alarming trends amongst America's youth and young adults. He cites numerous statistics which reveal an in- creasing number of young Americans are dropping out of high school or college, literacy rates are falling to greater depths each year,. The dumbest generation mark bauerlein pdf converter. The Dumbest Generation: How the Digital Age Stupefies You. And over one million other books are available for Amazon Kindle. It wouldn’t be going too far to call this book the Why Johnny Can’t Read for the digital age. Start reading The Dumbest Generation on your Kindle in.
Three individuals had a normal echocardiogram, but an abnormal ECG (IV-8, III-25, IV-11).Genetic investigation of family H640. Linkage analysis of recognized HCM genes was performed, and TNNI3 was identified as the likely disease gene. Mutation analysis of TNNI3 by direct sequencing identified a 87A→G nucleotide substitution of exon 8 resulting in an Asp190His amino acid substitution that segregated with the disease in the family (maximal two-point lod score: 4.8).Clinical and genetic investigation of other RCM patients. Following the identification of a disease causing TNNI3 mutation within family H640, investigation of the gene was performed in an additional nine unrelated patients who had been diagnosed with idiopathic RCM. The following six patients were shown to carry TNNI3 mutations.H417 was diagnosed at the age of 16 following a stroke associated with an episode of paroxysmal atrial fibrillation.
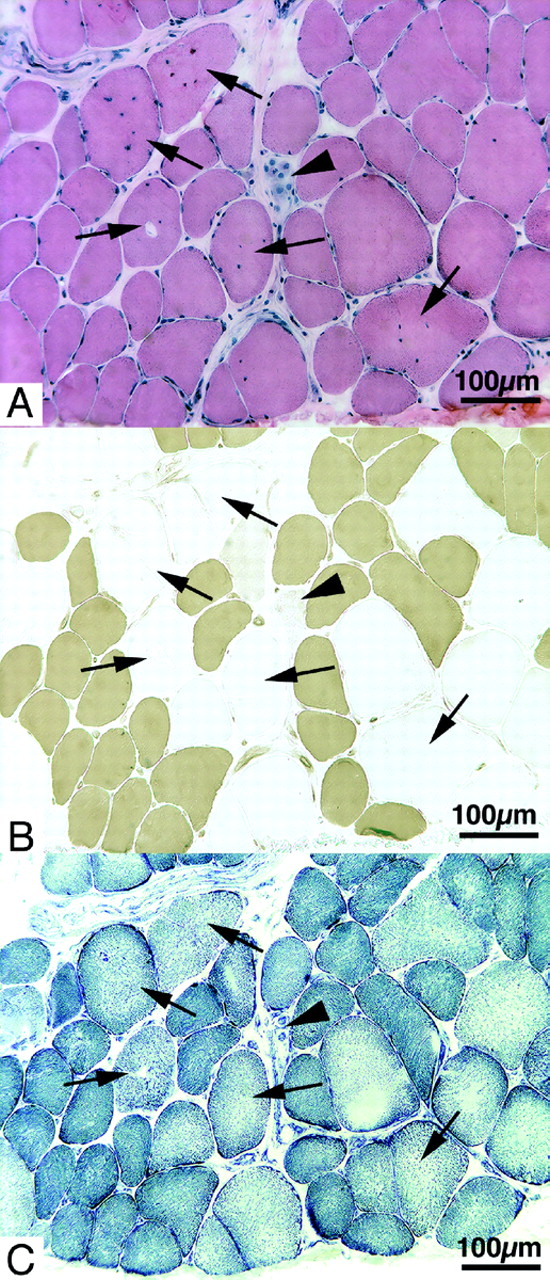
Echocardiography showed severe bi-atrial enlargement accompanied by normal left ventricular wall thickness and normal systolic function (Table and Figure a). ECG showed abnormal P-waves in all leads (Figure b). He died of heart failure at the age of 19 awaiting cardiac transplantation. Microscopy of right ventricular (RV) endomyocardial biopsies obtained antemortem showed nonspecific interstitial fibrosis without myocyte hypertrophy or myocyte disarray. Postmortem micro-scopy, however, revealed myofiber disarray, myocyte hypertrophy, and fibrosis similar to the findings in HCM patients (Figure c) (, ). Clinical features of restrictive cardiomyopathy in individual H417 affected by a de novo mutation in the gene for cardiac troponin I (Arg192His).
( a) Apical four-chamber echocardiogram in systole of H417 at the age of 18 with marked bi-atrial dilatation (left atrial LA size, 64 mm), dilatation of pulmonary veins (PV), normal-sized ventricles and normal wall thickness. LV, left ventricle; RV, right ventricle; RA, right atrium. ( b) 12-lead ECG in sinus rhythm with prominent P-waves in all leads, T-wave inversions, and incomplete right bundle branch block. ( c) Microscopy of heart tissue obtained postmortem with myocyte hypertrophy, abundant interstitial fibrosis, and myofibril disarray equivalent to the histological findings in HCM patients (, ) (hematoxylin and eosin staining, ×40).Direct sequencing of TNNI3 identified a 92C→A nucleotide substitution of exon 8, which resulted in an Arg192His amino acid substitution. The mutation was not present in either of his parents. Paternity was confirmed following haplotype analysis of 13 micro-satellite markers and the mutation was therefore considered to be a de novo mutation.H906 had failed to thrive since the age of 3 years and had a body weight of only 15 kg when idiopathic RCM was diagnosed at the age of 6 years. She had suffered from recurrent chest infections and frequent episodes of abdominal pain.
Her walking distance was less than 100 m. Echocardiography and ECG recordings showed typical features of RCM (Table ). Microscopy of RV endomyocardial biopsies showed nonspecific interstitial fibrosis as in biopsies from patient H417. The patient is currently awaiting cardiac transplantation. Mutation analysis of TNNI3 identified an 886A→G nucleotide substitution of exon 7, which resulted in a Lys178Glu amino acid substitution. The mutation was not present in her parents, and paternity was confirmed following haplotype analysis.
This mutation was also considered to be a de novo mutation.H805, H816, and H974 had typical features of RCM diagnosed in their late fifties (Table, and Figure ). H805 and H816 had symptoms of heart failure, while H974 was diagnosed following an embolic stroke. Micro-scopy of RV endomyocardial biopsies from H974 showed interstitial fibrosis with mild myocyte hypertrophy, but no myocyte or myofibrillar disarray. Mutation analysis of TNNI3 in both H805 and H816 identified the same 799C→T nucleotide substitution of exon 7 that lead to an Arg145Trp amino acid substitution. Haplotype analysis with respect to the TNNI3 locus on chromosome 19 did not suggest a common founder effect of the mutation. Their offspring were not carriers of the mutation, while remaining relatives were unwilling to participate in the study at present. Mutation analysis of H974 identified an 856G→A nucleotide substitution of exon 7 that led to an Ala171Thr amino acid substitution.
He was an only child, and no clinical data or DNA was available on his deceased parents. Subsequent mutation analysis of his children did not reveal further carriers.H38 developed symptoms of heart failure at the age of 17 years. Her echocardiogram was typical of RCM, except for a very localized mid-septal bulge (13 mm) without outflow tract obstruction. She died of heart failure at the age of 31 awaiting cardiac transplantation.
Postmortem microscopy of heart tissue showed myocyte and myofibrillar disarray, myocyte hypertrophy, and fibrosis similar to the findings in HCM patients (, ). Mutation analysis of TNNI3 identified a 797T→A nucleotide substitution of exon 7, which led to a Leu144Gln amino acid substitution.
Several members of her family were known to have died suddenly, but lived abroad and were not available for investigation.Mutation analysis. Prior to the genetic era of cardiomyopathies beginning with the identification of the first disease gene in HCM in 1990, it was believed that HCM, RCM, and the inherited form of dilated cardiomyopathy (DCM) were separate and distinct clinical and pathophysiological entities (, ). Much has been learned over the past decade, and it has been shown that sarcomeric gene mutations may be associated with both HCM and DCM ( ). These findings suggest that variation in the location of mutations within specific functional domains of sarcomeric disease genes can result in different clinical phenotypes (, ). The results of the current investigation show that RCM may also be part of the spectrum of hereditary sarcomeric contractile protein disease.The clinical expression of HCM in family H640 was unusual. The majority of affected HCM individuals had enlarged atria and/or evidence of restrictive filling with shortened DT in the presence of only mild to moderate left ventricle hypertrophy and normal cavity dimensions. Three gene carriers from family H640 had symptomatic clinical presentation that was dominated by restrictive filling pattern, marked atrial enlargement, and reduced left ventricular end-diastolic dimension (Table ).
Individual IV-10 had normal ventricular wall thickness, while individual IV-3 and III-17 had a small localized area of thickening at the apex. The phenotype of IV-3 and III-17 was characterized by RCM, and the severity of the changes was not readily explained by the small area of localized apical thickening (Table ). The number of premature sudden deaths in the family suggests that the Asp190His amino acid substitution identified is associated with an adverse phenotype. The mixed appearance of HCM and RCM in H640 have previously only been reported in three small families in which no genetic investigations were performed (, ). Previous studies have identified TNNI3 mutations in a small number of HCM families with insufficient phenotype information to determine whether these mutations were associated with a specific disease expression ( – ).The finding of three patients in family H640 with a phenotype dominated by restrictive physiology lead us to hypothesize that idiopathic RCM could be part of the clinical expression of TNNI3 mutations. Another nine patients fulfilling RCM diagnostic criteria were investigated for mutations in TNNI3. Their clinical presentation was with embolic stroke, heart failure, or for transplant evaluation.
They had severe restrictive filling on Doppler, marked left atrial dilatation, and low-normal or reduced left ventricular end-diastolic dimension with normal left ventricular wall thickness (Table ). Two young individuals (H417, H906) had severe and early onset of disease consistent with them being carriers of de novo mutations. The histology of RV endomyocardial biopsies from three patients (H417, H906, H974) showed nonspecific interstitial fibrosis without myocyte or myofiber disarray. Postmortem microscopy of cardiac tissue from two patients (H906, H38), however, showed typical HCM histology (Figure b). These apparent differences may reflect difficulties in obtaining representative myocardial tissue by biopsy as reported previously ( ). The finding of typical HCM histology when the whole heart was available for evaluation confirms previous observations of hearts from patients with a clinical diagnosis of RCM ( ).The genetic investigations of the current study identified six novel TNNI3 mutations, which were all localized in conserved and functional important regions of the gene (Table and Figure ).
It is well established that regulation of muscle contraction is primarily dependent on the intracellular concentration of Ca 2+. The major sensor of the intracellular Ca 2+ level is the troponin complex, which is composed of three subunits, troponin C, troponin I, and troponin T. All three troponins are located within the thin muscle filament, and their primary function is to control the interaction between the thick and thin filament during muscle contraction and relaxation. Troponin I can bind to actin-tropomyosin and prevent muscle contraction by inhibition of actin-tropomyosin–activated myosin (actomyosin) ATPase activity. The inhibitory effect of troponin I is reversed by troponin C following binding of Ca 2+, which subsequently introduces conformational changes in the entire troponin complex leading to muscle contraction ( ). Previous studies have defined different functional domains of TNNI3 (Figure ) ( ). It has been shown that the sequence required for inhibition of human cardiac troponin I actomyosin ATPase activity consists of 21 amino acid residues from number 137 to 148 ( ).
Two of the RCM mutations identified, Leu144Gln and Arg145Trp, are localized within this important region of TNNI3. The remaining RCM mutations are localized in proximity to or within other important domains of troponin I necessary for the normal function of the troponin complex. The Ala171Thr and Lys178Glu mutations may influence the inhibitory function through actin binding since previous studies have shown that amino acids number 173–181 bind to actin and increase the inhibitory effect of troponin I. The Asp190His and Arg192His mutations are both localized within the conserved C-terminal region of the protein, which is also required for normal inhibitory function of troponin I.Several recent in vitro expression studies have investigated previous reported TNNI3 mutations associated with HCM ( – ). These HCM mutations were found to be associated with increased Ca 2+ sensitivity and diminished inhibition of the actomyosin ATPase activity. Similar findings were reported following analysis of transgenic mice expressing an Arg145Gly HCM mutation ( ). The hearts of transgenic mice showed increased contractility, whereas relaxation was impaired.The diversity of the phenotypic expression of TNNI3 mutations in families with a mixed appearance of HCM and RCM is not explained by the current knowledge of the molecular and functional impact of TNNI3 mutations and suggest that additional genetic and environmental factors influence disease expression.
However, the finding of TNNI3 mutations in a substantial proportion of patients diagnosed with idiopathic RCM suggest a causal relationship between gene abnormality and disease. This is further strengthened by the fact that de novo mutations were identified in patients with severe and early onset RCM. Previous expression studies of TNNI3 mutations in HCM reveal alterations in Ca 2+ sensitivity and inhibitory function that may also influence the development of RCM.
The identification of TNNI3 mutations in RCM patients may serve as a model for the development of a similar phenotype in transgenic animals to further investigate pathophysiology and diastolic abnormalities (filling, relaxation, compliance) in the presence of normal systolic function and wall thickness.In summary, in patients with idiopathic RCM evaluation of the family or histology from the whole heart may reveal clinical or pathological features of HCM. The fact that TNNI3 mutations were identified in a significant proportion of such patients indicates that idiopathic RCM is part of the clinical expression of sarcomeric contractile protein disease and of HCM. Troponin I disease should be considered in patients with apparent idiopathic RCM and family investigation for both restrictive and hypertrophic cardiomyopathy should be offered to relatives of these patients. Free wifi password templates.
In 2011, the Institute of Medicine (now the National Academy of Medicine) published Relieving Pain in America: A Blueprint for Transforming Prevention, Care, Education, and Research. This landmark report heralded a new era for pain relief, one in which the public health impact of limitations in providing effective pain relief would be pushed to society's forefront. The timing of the Relieving Pain in America report seems especially prescient given the fallout from the ongoing opioid crisis.Although the public health implications of Relieving Pain in America clearly resonates, another important part of the report is the call for a “transformed understanding of pain.” A quote from the preface nicely summarizes what a transformed understanding may entail:Our committee recognizes the need for a transformed understanding of pain. We believe pain arises in the nervous system but represents a complex and evolving interplay of biological, behavioral, environmental, and societal factors that go beyond simple explanation”The purpose of this Point of View is to review foundational evidence that supports pain as a nervous system condition. The goal is to increase awareness among musculoskeletal-focused physical therapists that making meaningful progress in improving pain management is likely to involve integration of nervous system–related factors. We highlight plasticity and automaticity as specific examples with high potential to change entrenched education, practice, and research approaches in such a way that our understanding of musculoskeletal pain continues to improve. Chronic Pain as a Nervous System DisorderRelieving Pain in America’s assertion that “pain arises in the nervous system” is not surprising.
There has always been a strong central nervous system (CNS) component for pain perception (eg, the Gate and Neuromatrix theories). However, what has substantially changed is that the development of chronic pain, which was once thought to be contingent on peripherally driven input into the nervous system, is now believed to be mitigated by changes inherent to the nervous system.
This shift in emphasis has resulted in a more complex model in which CNS parameters dominate the clinical phenotype of chronic pain, but the phenotype still has relevant peripheral components.There is sufficient evidence to support this assertion, and it is beyond the scope of this Point of View to provide a comprehensive review. Suffice to say that, given our current knowledge base, it is clear that functional and structural changes in the CNS are associated with many chronic pain syndromes of different etiologies, such as chronic low back pain (LBP), fibromyalgia, pelvic pain syndrome, spinal cord injury, and phantom limb pain. The normal state of the nervous system is a balance between automaticity (brainstem and spinal cord, represented in yellow) and executive control (cortex, represented in red). Efficient and effective performance of both gait and pain modulation is dominated by automaticity (indicated by the larger area of yellow shading)–processing that is fast, in parallel, and requires little effort even with increasing workload.
After insult and changes to the nervous system (ie, damaged state), control of both gait and endogenous pain modulation becomes dominated by executive control (indicated by the larger area of red shading). Then, the contrast between intervention target strategies used is stark (indicated by differences in shading between stroke and chronic pain). The majority of the focus in gait rehabilitation is on regaining automaticity. Rehabilitation in chronic pain often emphasizes executive control and includes exogenous inhibitory agents rather than building endogenous capacity. Despite the differences in intervention target and management strategies, the ideal outcome remains the same: the resumption of dominance in automaticity, demonstrated by efficient and effective gait and pain modulation (indicated by the similarities in shading between stroke and chronic pain). The normal state of the nervous system is a balance between automaticity (brainstem and spinal cord, represented in yellow) and executive control (cortex, represented in red).
Efficient and effective performance of both gait and pain modulation is dominated by automaticity (indicated by the larger area of yellow shading)–processing that is fast, in parallel, and requires little effort even with increasing workload. After insult and changes to the nervous system (ie, damaged state), control of both gait and endogenous pain modulation becomes dominated by executive control (indicated by the larger area of red shading). Then, the contrast between intervention target strategies used is stark (indicated by differences in shading between stroke and chronic pain). The majority of the focus in gait rehabilitation is on regaining automaticity.
Rehabilitation in chronic pain often emphasizes executive control and includes exogenous inhibitory agents rather than building endogenous capacity. Despite the differences in intervention target and management strategies, the ideal outcome remains the same: the resumption of dominance in automaticity, demonstrated by efficient and effective gait and pain modulation (indicated by the similarities in shading between stroke and chronic pain).Assuming the reader accepts these parallels, there are future implications to consider for improving pain management. The first implication is that existing therapies may provide pain relief through restoring automaticity for pain inhibition.
Consider muscle- or joint-based manual therapy, as they have an established inhibitory effect on nervous system processing of nociceptive input., The primary way manual therapy was hypothesized to provide pain relief was through inhibition of facilitatory input. Evidence supporting the importance of endogenous pain modulation, however, was observed in patients who were healthy receiving spinal manipulation and patients with knee osteoarthritis receiving joint mobilization. For therapies such as manual therapy, a mechanistic shift of action from inhibition to enhanced modulation of nociception will not directly impact efficacy, but it could be a theoretical foundation for creating next-generation delivery approaches.The second implication is whether there is value in training automaticity in pain relief by intentionally having an individual experience pain without harm. Training automaticity via locomotor training has shown equivalent benefit to physical therapist–supervised home exercise for walking recovery, but similar paradigms testing the efficacy of automaticity for treating pain through the experience of pain have not been developed. Instead, most pain management approaches rely on pharmacological or nonpharmacological agents to inhibit pain. Paradoxically, training automaticity for pain relief would provoke pain, under the assumption that this provocation would build psychological and biological resilience over time. This may sound counter to a “do no harm” approach to patient care, but precedent exists from a systematic review outlining the short-term advantage of experiencing pain with exercise for reducing pain intensity.
Much more research must be completed in this area before it is established when experiencing pain may be beneficial by developing resiliency.The third implication is to avoid solely relying on executive control for providing pain relief. In routine care episodes, there appears to be some advantage to incorporating approaches that increase cognitive load (eg, neuroscience education, cognitive behavioral therapy, and psychologically informed approaches).
These interventions all involve a level of executive control necessary in developing adaptive pain beliefs and behaviors; however, their use comes at a cost of limiting the potential for automaticity. The overall balance between executive control and automaticity in a treatment plan may be especially important to consider if full recovery is still a feasible treatment goal. Think again of enhancing motor recovery following stroke. These treatment paradigms involve training that emphasizes automaticity by performing the task directly, with repetition and variation., Executive control is used as an adjunct to the automaticity-focused approach by providing feedback that allows for motor learning. As best we can tell, there are very few programs that are focused entirely on education and cognitive behavioral therapies to enhance walking; instead, patients are given the opportunity to enhance recovery by performing the task. Perhaps it is time to take a similar stance in promoting automaticity for pain relief when developing the next generation of nonpharmacological treatments for musculoskeletal conditions.
ConclusionsThis Point of View focused on the roles that plasticity and automaticity could play when chronic musculoskeletal pain is conceptualized as a nervous system disorder. The assertions we make are speculative and represent an attempt at generating hypotheses and providing direction for integrating research in areas not typically integrated (ie, it is rare to have a combination of nociceptive, motor control, and movement sciences ).A shift from focusing on peripheral input to acknowledging the importance of the CNS is not meant to encourage mutual exclusiveness. Both systems are relevant to the development of chronic pain syndromes, with special interest in how the peripheral and central systems interact to perpetuate or extinguish nociceptive signals.
The inherent complexity of these interactions has yet to be captured in a way that enhances clinical decision making. The available research suggests that conditions characterized as central sensitization syndromes also exhibit signs of peripheral sensitization (and vice versa). For example, in our own work involving patients with unilateral shoulder pain scheduled for arthroscopic surgery, we found evidence for local and remote pain sensitivity to exist in all 4 possible combinations of proxy measures (ie, peripheral sensitivity only, peripheral and central sensitivity, central sensitivity only, or no evidence of sensitivity).The temporal association of peripheral and central systems is another layer of complexity to consider. It is often assumed that the ongoing peripheral input leads to the development of central sensitization.
However, in our aforementioned study in shoulder pain, there was evidence of elevated remote pain sensitivity with and without elevated local pain sensitivity. These examples indicate that more work is needed before we fully understand the spatial and temporal interplay between peripheral and central systems involved with chronic pain development. There are likely to be both peripheral and central treatment targets, along with optimal times for their application, for providing effective pain relief; however, we haven’t identified these windows of opportunity yet.The assertions in this Point of View may spur physical therapists to explore management models that go beyond only providing pain relief (eg, movement as a way to alter the relevance of pain to the individual beyond only a reduction in pain intensity ). We fully acknowledge that more scientific work is to be done before speculations such as these are actualized into gains in patient care; however, we also acknowledge there is a good chance that paradigm-shifting improvements in patient care may emerge through better integration of what we know about providing pain relief and what we know about the nervous system. Look for PTJ’s Special Issue on the Nonpharmacological Management of Pain in May Author ContributionsConcept/idea/research design: Steven Z. George, Mark D. BishopWriting: Steven Z.
George, Mark D. BishopConsultation (including review of manuscript before submitting): Steven Z. George FundingS.Z. George's and M.D. Bishop's writing time was supported through a grant from the National Institutes of Health/National Institute of Arthritis and Musculoskeletal and Skin Diseases (ref. DisclosuresThe authors completed the ICJME Form for Disclosure of Potential Conflicts of Interest and reported no conflicts of interest.
. d. d.
d. d. d. d. d. d. d.
d. d. d.
d. d. d. d. d.
d. d. d. d. d. d.
d. d.
d. d. d. d.
d. d.
d. d. d. d.
d. d. d. d. d. d. d.
d. d. d. d. d. d. d.
d. d. d. d. d. d.
d. d. d. d. d. d. d.
d. d. d. d. d. d. d.
d. d. d. d. d.
d. d. d. d. d. d.
d. d. d. While the following list is not intended to be comprehensive, it represents many of the tools commonly used in rehabilitation assessment of patients with MS.Ashworth andThese are ordinal scales of tone intensity. The Ashworth rates tone on a scale of 0–4, while the Modified Ashworth was developed to further define the lower end of the scale making it more discrete by adding the grade 1+ (Lee KC, Carson L, Kinnin E, and Patterson V. The Ashworth Scale: A reliable and reproducible method of measuring spasticity.
Comments are closed.
|
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
- Blog
- Blog
- Blog
- Blog
- Blog
- Blog
- Blog
- Blog
- Blog
- Blog
- Blog
- Blog
- Blog
- Blog
- Home
- Bosch Wtl 5200 Manual Transfer
- Free Download Mafia 1 Pc
- Adobe Indesign Cs5 Keygen Download Free
- Download Free Project Altered Beast Isolation
- Max Payne 2 Blood Models
- Download Resume In Html Format Online
- Uno Nessuno Centomila Ebooking
- Dazzle Fusion Drivers Windows 7
- New Super Mario Bros 3 Rom Hack - Download Free Apps
- Ofra Haza Desert Winrar
- Download Avast
- Rhinoceros 5 Beta Keygen Software
- Free Spirit Treadmill Owners Manual
- The Dumbest Generation Mark Bauerlein Pdf Converter
- Power Plant Performance By Gill Pdf Merge
- Download Wifi Splash Page Template Free Software
- Edirol Ua-5 Driver Windows 10
- Sissy That Walk Download Mp3
- Drivers Placa Mãe Ecs P4vmm2
- Les Miserables Piano Pdf Scribd Review
- Best Video Editor Free Download Windows 7
 RSS Feed
RSS Feed
